Agnieszka Lipinska
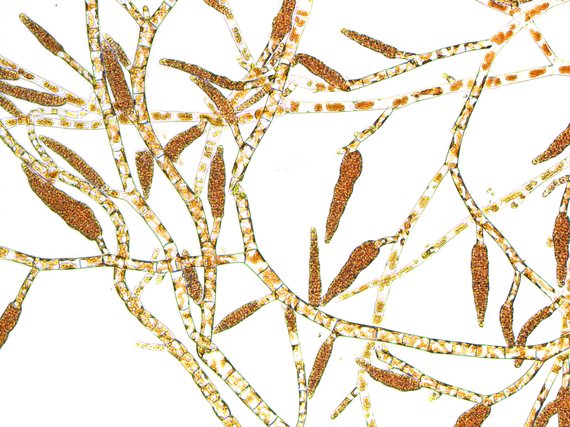
Reproductive isolation and speciation in the brown algae
Max Planck Institute for Biology Tübingen
Adjunct faculty in: IMPRS
Vita

- PhD, Ghent University, Belgium (2013)
- Postdoctoral fellow, Station Biologique de Roscoff, CNRS-Sorbonne University, CNRS, France (2013-2018)
- Junior Group Leader, Algal Genetics Group, Station Biologique de Roscoff, CNRS-Sorbonne University, Roscoff, France (2018-2020)
- Group Leader, Department of Algal Development and Evolution, MPI for Biology, Tübingen, Germany (since 2020)
Research Interest
Reproductive barriers are important because they prevent gene exchange between diverging species, helping to maintain species integrity. Therefore, the identification of genomic regions responsible for the evolution of reproductive isolation has been a major focus in speciation genetics. Numerous reproductive barriers evolve during the speciation process, including pre-mating and post-mating incompatibilities between diverging species. Post-mating barriers are often attributed to negative epistatic interactions between genetic elements in the hybrid offspring and empirical evidence suggests that sex chromosomes play an important role in this interplay.
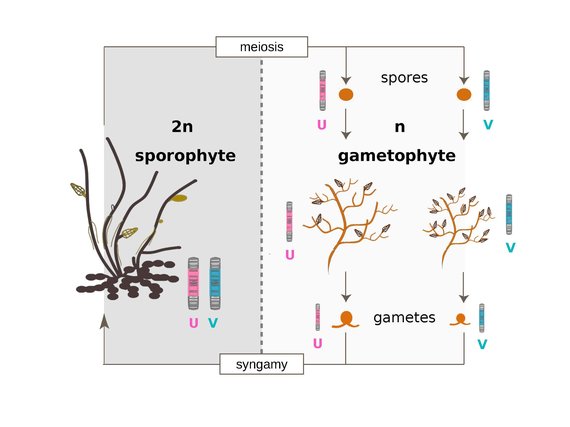
To date, many studies have investigated the genomic architecture of speciation, and particularly the role of sex chromosomes, using a handful of organisms in the plant and animal kingdoms. Our goal is to gain insight into the genetic bases of reproductive isolation using a novel, brown algal model (Ectocarpus) that is extremely distantly related to both animals and green plants. We aim to characterize the role of sex chromosomes in postzygotic isolation, to identify the causative genomic regions and to carry out evolutionary analyses of the genes underlying hybrid incompatibility. Moreover, Ectocarpus, with its haploid-diploid life cycles and a UV sex determination (fundamentally different from XY and ZW systems), is expected to provide important new insights into mechanisms of speciation and the universality of the underlying processes across major eukaryotic supergroups.
Available PhD Projects
- Currently not recruiting PhD students
Selected Reading
- Lipinska AP, Toda NRT, Heesch S, Peters AF, Cock JM, Coelho SM. 2017. Multiple gene movements into and out of haploid sex chromosomes. Genome Biology 18: 104.
- Lipinska AP, Van Damme EJM, De Clerck O. 2016. Molecular evolution of candidate male reproductive genes in the brown algal model Ectocarpus. BMC Evolutionary Biology 16: 5.
- Avia K, Lipinska AP, Mignerot L, Montecinos AE, Jamy M, Ahmed S, Valero M, Peters AF, Cock JM, Roze D, et al. 2018. Genetic Diversity in the UV Sex Chromosomes of the Brown Alga Ectocarpus. Genes 9: 286.
- Hatchett WJ, Jueterbock A, Kopp M, Coyer JA, Coelho SM, Hoarau G, Lipinska AP. 2022. Evolutionary dynamics of sex-biased gene expression in a young XY system: Insights from the brown algae. (in press, New Phytologist)
- Lipinska A, Cormier A, Luthringer R, Peters AF, Corre E, Gachon CMM, Cock JM, Coelho SM. 2015. Sexual Dimorphism and the Evolution of Sex-Biased Gene Expression in the Brown Alga Ectocarpus. Molecular Biology and Evolution: msv049.